PRGMiner Tutorial
A comprehensive guide to using PRGMiner for plant resistance gene prediction
Homepage
The homepage provides an overview of PRGMiner and quick access to all major features.
- Navigation bar with links to all major sections
- Quick introduction to PRGMiner's capabilities
- Direct access to prediction tools
- Latest updates and announcements
Making Predictions
Learn how to submit sequences and get predictions using PRGMiner.
Input Methods
Accession ID
Enter a valid protein accession ID from NCBI or UniProt to fetch and analyze the sequence.
FASTA File
Upload a FASTA file containing one or multiple protein sequences for analysis.
Paste Sequence
Directly paste FASTA-formatted sequences into the text area.
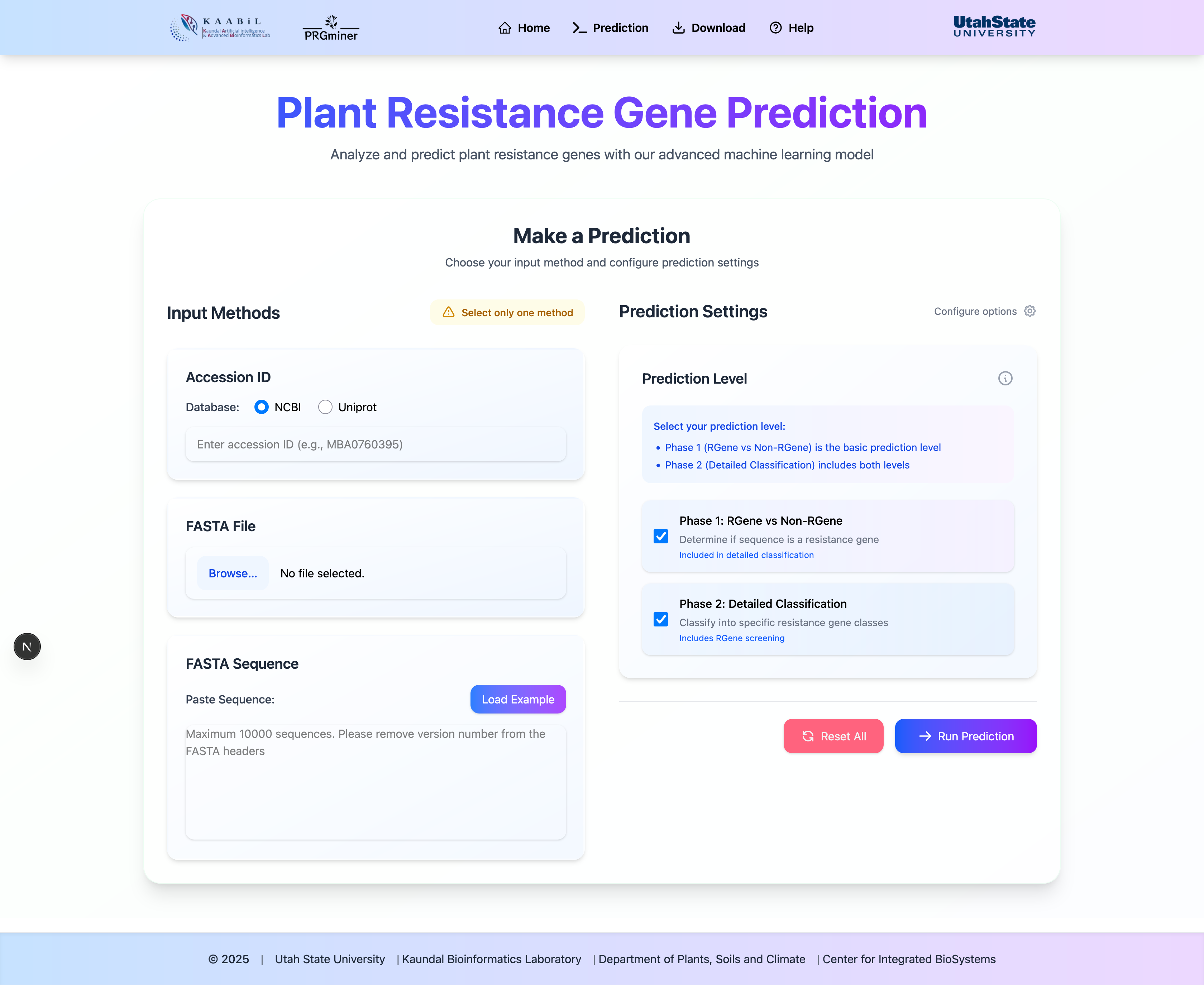
Submission Process
- Choose your preferred input method
- Click "Run Prediction" to start the analysis
- Wait for the analysis to complete
Understanding Results
Learn how to interpret and download your prediction results.
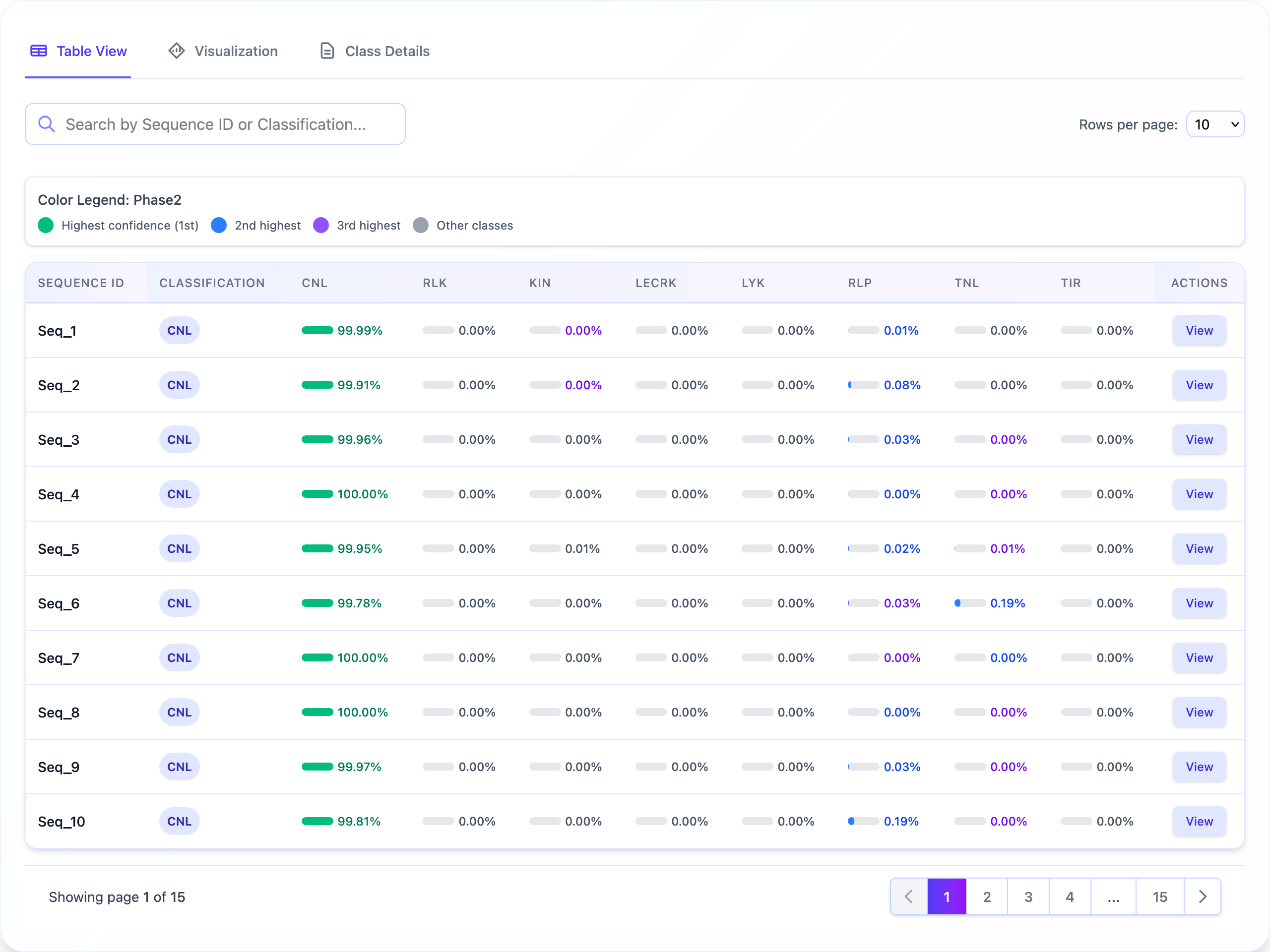
Results Table
- Sequence ID and basic information
- Prediction outcome (R-gene or Non-R-gene)
- Confidence scores for predictions
- Detailed classification for R-genes
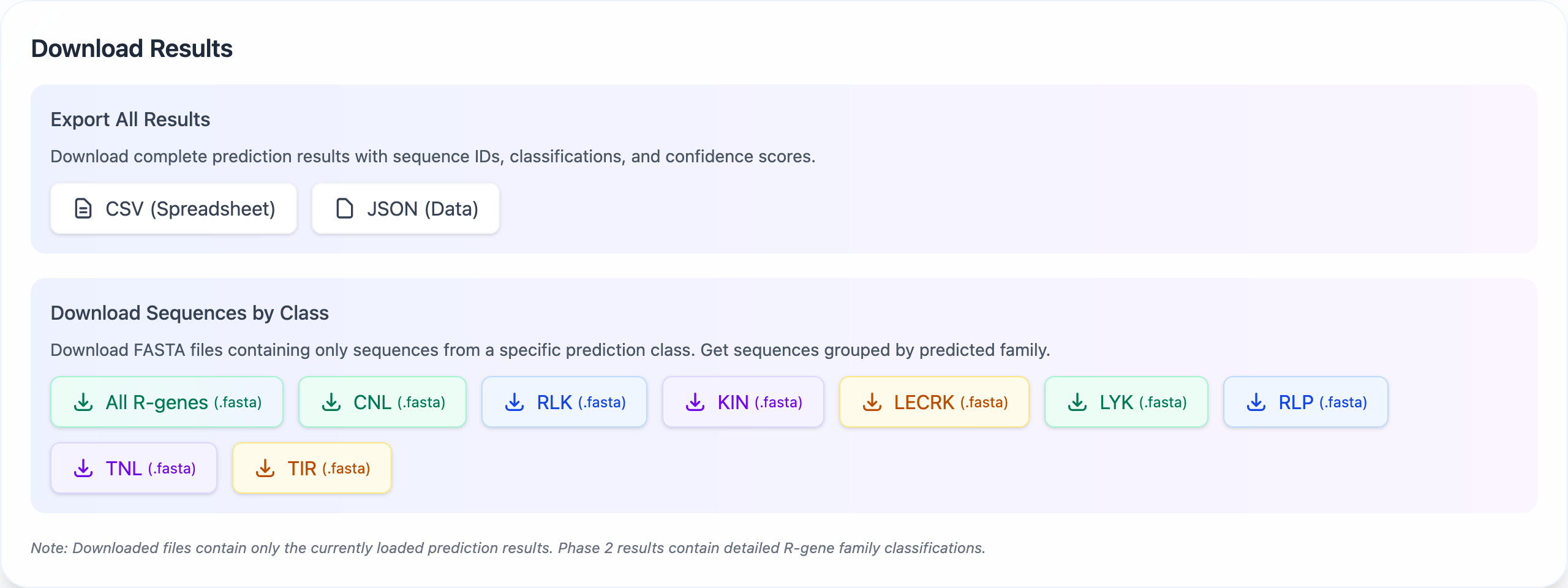
Download Options
Complete Results
Download all results in CSV, JSON, or FASTA format, including sequences and predictions.
Filtered Results
Download results for specific R-gene classes or confidence thresholds.
Documentation and Help
Access comprehensive documentation and get help when needed.
Documentation
Detailed technical documentation covering:
- Installation guide
- API reference
- File format specifications
- Best practices
Help Center
Get assistance through:
- FAQs
- Email support
- GitHub issues
- Community forums
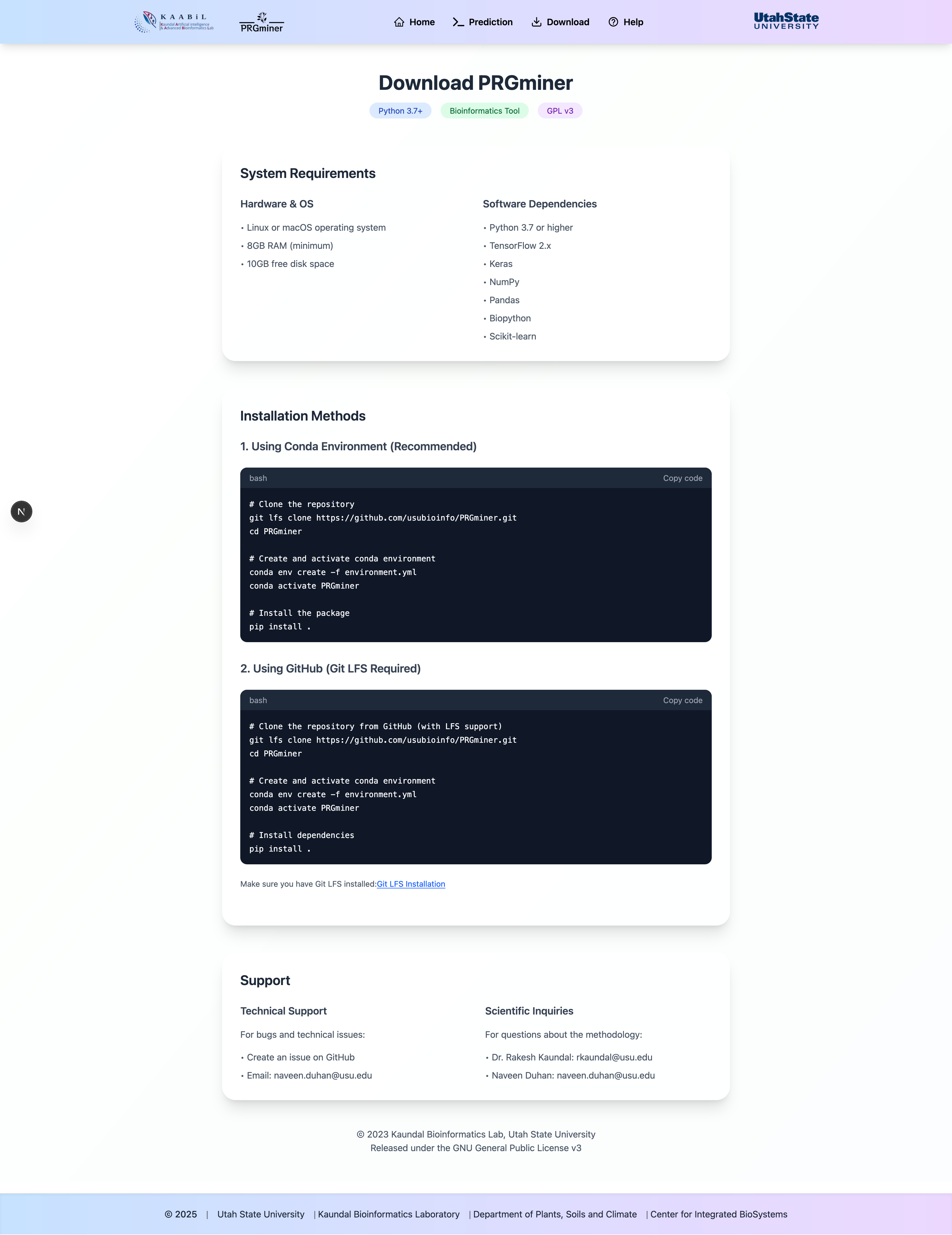
Local Installation
Instructions for downloading and running PRGMiner locally.
System Requirements
- Python 3.7 or higher
- Required dependencies (listed in requirements.txt)
- Sufficient RAM for large datasets
- GPU support (optional, for faster processing)
Important Note
Local installation is recommended for:
- Processing large datasets (>10,000 sequences)
- Integration with existing pipelines
- Customized analysis workflows
- Offline usage